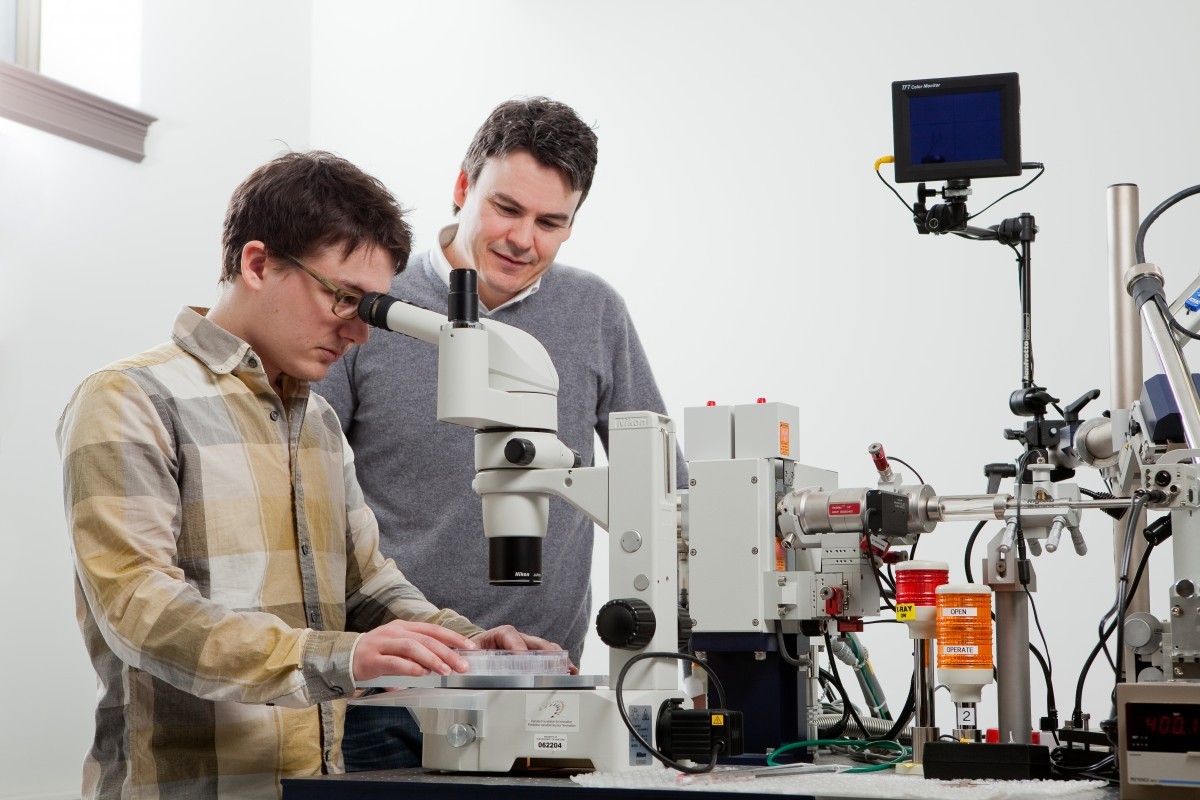
PhD student Ben Bailey-Elkin locates a protein crystal under the microscope
New twist in an ancient fight
Removing bacteria’s defence gives us a new offence
We can’t win, so we’re trying not to lose. Since the late 1980s we’ve struggled to find or build new weapons to use in our ancient fight against bacteria and now bacteria are developing resistance at an alarming rate to the drugs we have.
Global health agencies publish a frightening number: in 35 years, if left unchecked, antibiotic-resistant bacteria will kill 10 million people annually across the globe. For those keeping score, that’s two million more than cancer.
One microscopic foe is Pseudomonas aeruginosa. It evolved in soil, which looks benign to our eye but is actually a surprising bed of microscopic violence and struggle. For 3.5 billion years bacteria have fought to survive here, developing new defences and weapons every generation, which for bacteria is about every 20 minutes.
As a result, bacteria today are a hearty bunch – Pseudomonas notoriously so. It grew up in this harsh underworld but now lives a relaxing life in hospitals, infecting lungs and open wounds.
Its only combatant is us, we have only a few drugs to fight it, and they are quickly losing potency.
“How is it doing this? How is it defending itself from our drugs? That’s what we’ve been trying to figure out and a key clue comes from the materials it produces,” says Brian Mark, an associate professor of microbiology at the University of Manitoba who has dedicated a branch of his research program to fighting this bacterium.
Pseudomonas creates a few materials, one of which is a nearly impenetrable outer membrane.
Most drugs can’t get through it, and if any do, Pseudomonas’ powerful efflux pumps quickly toss them out. One type of drug that can defeat these systems is beta-lactam. But once inside, Pseudomonas detects it and quickly produces a protein to destroy the drug.
At this juncture, we have two options. Develop new antibiotics, like U of M microbiologist George Zhanel and U of M chemistry and medical microbiologist professor Frank Schweizer have been doing since 2006. Or, study the materials Pseudomonas uses to defend itself to see how we can help existing antibiotics fight better.
Mark’s lab studies the structure of defensive proteins Pseudomonas creates to find their exploitable weaknesses. His team begins by extracting the proteins and coaxing them to crystallize. Once the protein crystal is sufficiently large, it is brought to an X-ray diffraction machine, which bombards it with X-rays to reveal its atomic structure.
Every protein is built out of a set of 20 different amino acids combined into long and unique chains. Then, crucially, it folds into a specific shape and that shape dictates its function. So when Mark investigates what the protein looks like three-dimensionally, he gains insight into how it works.
“There are always secrets that pop up,” Mark says. “When you solve the 3-D structure of a protein you’ve never seen before, there are always interesting surprises that you can see that you never expected. The solution Mother Nature has arrived at to carry out some function is sometimes completely different than we had theorized.
Either the way it folds or the way it interacts with another molecule can be completely different than what we expect. There are always surprises that show up.”

Mark in front of the X-ray diffractormeter that is used to investigate protein structure in atomic-scale detail
In the case of Pseudomonas, Mark examines the proteins it uses to activate the production of beta-lactamase, a protein that destroys many of our most cherished antibiotics known as beta-lactams. After solving the 3-D structure of these proteins, his team looks for areas on the protein where small, drug-like molecules can fit.
Once snug in place, the small-molecules designed by Mark and his collaborator professor David Vocadlo, a chemical biologist at Simon Fraser University, block Pseudomonas production of beta-lactamase. In short, they took its defence away and in the process showed beta-lactam resistance could be overcome in Pseudomonas using such a strategy.
This approach is called structure-based drug design: Solve the shape; find the weakness; exploit it. It works well, except the process has one hurdle – it requires scientists to crystallize the proteins they wish to target.
“Often times you put a lot of effort into all the steps and then you run into the problem that you can’t crystallize the protein you want to study.” Mark says. “You never know a priori the chemical conditions that will give rise to a protein crystal. This can be a stumbling block that is very hard to overcome.”
Much to the joy of Mark, one scientist recently opened a new avenue of opportunity in the fight against antibiotic drug resistance: Kim Lewis, from Northeastern University in Boston, who gave this year’s U of M’s Armes Lecture in the department of chemistry.
The vast majority of microbes require very specific conditions to grow. To discover new sources of antibiotics – weapons bacteria make to defend themselves from peers – requires us to grow them. But soil, the cradle we want to rob, has a colossal supply of niches for life to exploit. If a soil microbe slides micrometers in any direction, it may occupy a new niche – one with different acidity, moisture or chemical factors, that may not be habitable to it. We cannot replicate this great quantity of environments in the lab.

Thus, Lewis thought up the idea of culturing bacteria in the soil itself. Basically, you put the bacteria you want to culture into a cassette and insert it back into their environment. Whatever factor it needs to grow passes through the permeable cassette walls. The bacteria grow, you retrieve the cassette and study the culture.
“It’s brilliant,” Mark says. “And he’s now discovered a new antibiotic that works really well against a major class of bacteria. There is considerable potential to find new antibiotics from all of these uncultured microbes. It appears we’ve only scratched the surface.”
Microbiologists celebrate good news like this when they get it, but remain humble because history suggests they should. For instance, on May 15, 1944, as the Allied troops began preparing for D-Day, Time magazine published a picture of Dr. Alexander Fleming on the cover with the text: “His penicillin will save more lives than war can spend.” As that cover ran, however, researchers began noticing certain strains of staphylococci no longer responded to penicillin.
The next great drug discovered was methicillin in 1959.
It was revered, but 11 months later bacteria resistant to methicillin were discovered.
This story repeats over and over again: we find a drug, then we discover bacteria already have a defence against it, one likely developed in ancient times.
“And the resistance mechanisms can be shared among bacteria – this is the real problem,” Mark says. “Some harmless soil bacteria may have a remarkable defense
mechanism against an antibiotic we rely on, yet it doesn’t pose a medical risk because it’s not infectious. However, bacteria can share DNA with each other, so if an infectious microbe acquires the mechanism, and then shows up in the clinic, it can become a serious problem.”
This is why Mark’s approach offers such an exciting avenue: design materials to deactivate defences, allowing old drugs to work once again.
“Bacteria are smarter than us,” Zhanel quips. “They are working very hard to resist and develop resistance to what we are doing. So we are playing catch-up, and our goal is to continue playing catch-up with a variety of strategies.”
By understanding biological materials such as proteins and designing new drugs to block them we may indeed gain the upper hand in the battle against antibiotic drug resistance.
We may not win, but we won’t lose. “Keep in mind that we’re studying the material of Mother Nature,” Mark says.
“She’s the master of material science….We have a lot of challenges ahead.”