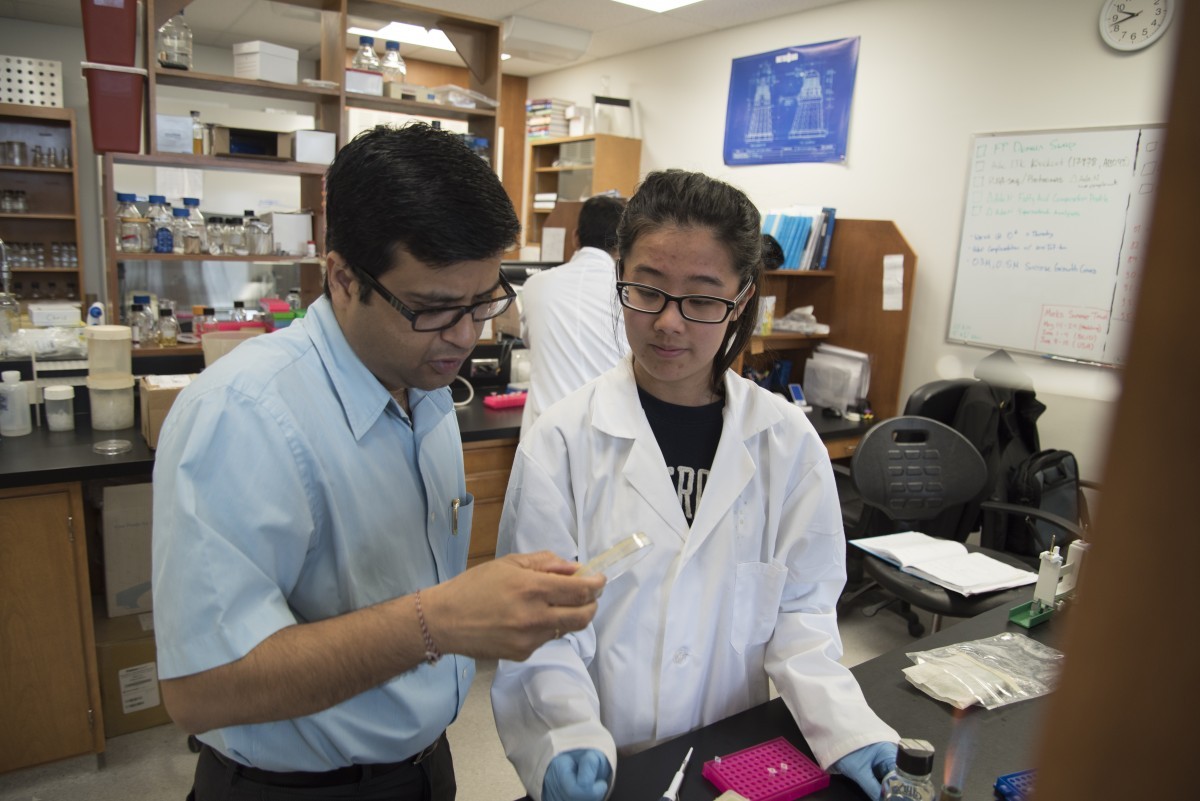
Ayush Kumar (Left) of Kumar Lab, of the Faculty of Science's Department of Microbiology
CIHR and Research Manitoba Funding for U of M Microbiology Labs
Three Microbiology Labs Receive Funding Towards Continuing Their Research
It was announced this week that two labs from Microbiology received funding from the Canadian Institutes of Health Research and one lab from Microbiology received funding from Research Manitoba. The funding ranges from $100,000 to $760,680 – over periods of 1 to 5 years.
This funding will allow our researchers to continue their work in the follow areas:
Kumar Lab
Pseudomonas aeruginosa is a bacterium notorious for causing various infections in immunocompromised individuals. Resistance of P. aeruginosa to multiple antibiotics makes the eradication of infections a challenge.
Although P. aeruginosa uses various mechanisms to resist the action of antibiotics, an energy-dependent outflow of antibiotics using so-called “RND-pumps” is one of the key mechanisms. Using this mechanism, P. aeruginosa transfers antibiotics out of the cell and therefore prevents antibiotic molecules from reaching an effective concentration.
The functional RND-pumps are formed by three different proteins; the goal is to understand the mechanisms by which these three proteins interact. Previous work from our (and our collaborator’s) laboratory shows that one of the three components of the RND-pumps that is present in the bacterial outer membrane may play a role in determining which antibiotic is recognized by the RND-pump. This novel finding may also lead towards new avenues of investigation that might lead to the inhibition of RND-pumps. Further, a number of state-of-the-art genetic tools for P. aeruginosa have been developed that will aid in studying the mechanisms of interaction between different components of the RND-pumps.
Mark Lab
The effectiveness of beta-lactam antibiotics is being eroded by the increasing prevalence of bacteria that produce AmpC beta-lactamase, an enzyme that inactivates most clinically available beta-lactam antibiotics. AmpC is produced by many pathogenic bacteria including Pseudomonas aeruginosa, a pathogen that is the leading cause of lung infections and mortality among people with cystic fibrosis.
Our objective is to fully understand the biochemical pathway that allows P. aeruginosa to induce production of AmpC in response to beta-lactam treatment and use this knowledge to guide new therapeutic approaches to block this mechanism of antibiotic resistance.
Pelka Lab
Adenoviruses are a family of small DNA viruses that have been shown to cause cancer in rodents. One of the adenovirus genes responsible for initiation of cancer is the E1A gene, which is the first gene expressed after cells are infected with the virus.
Due to its tendency to cause tumors, E1A is an ideal tool to study carcinogenesis, due to the fact that in early infection the virus mimics many of the same events that occur in cancer. We will exploit this property of E1A to investigate how E1A modulates the activities of RuvBL1, a protein that is frequently overexpressed in cancer and which functions in processes highly relevant to carcinogenesis. The proposed work aims to look for novel factors that modulate the cell cycle and differentiation, processes that are not properly regulated in cancer. Understanding these processes on a fundamental level is crucial to developing new cancer therapies.