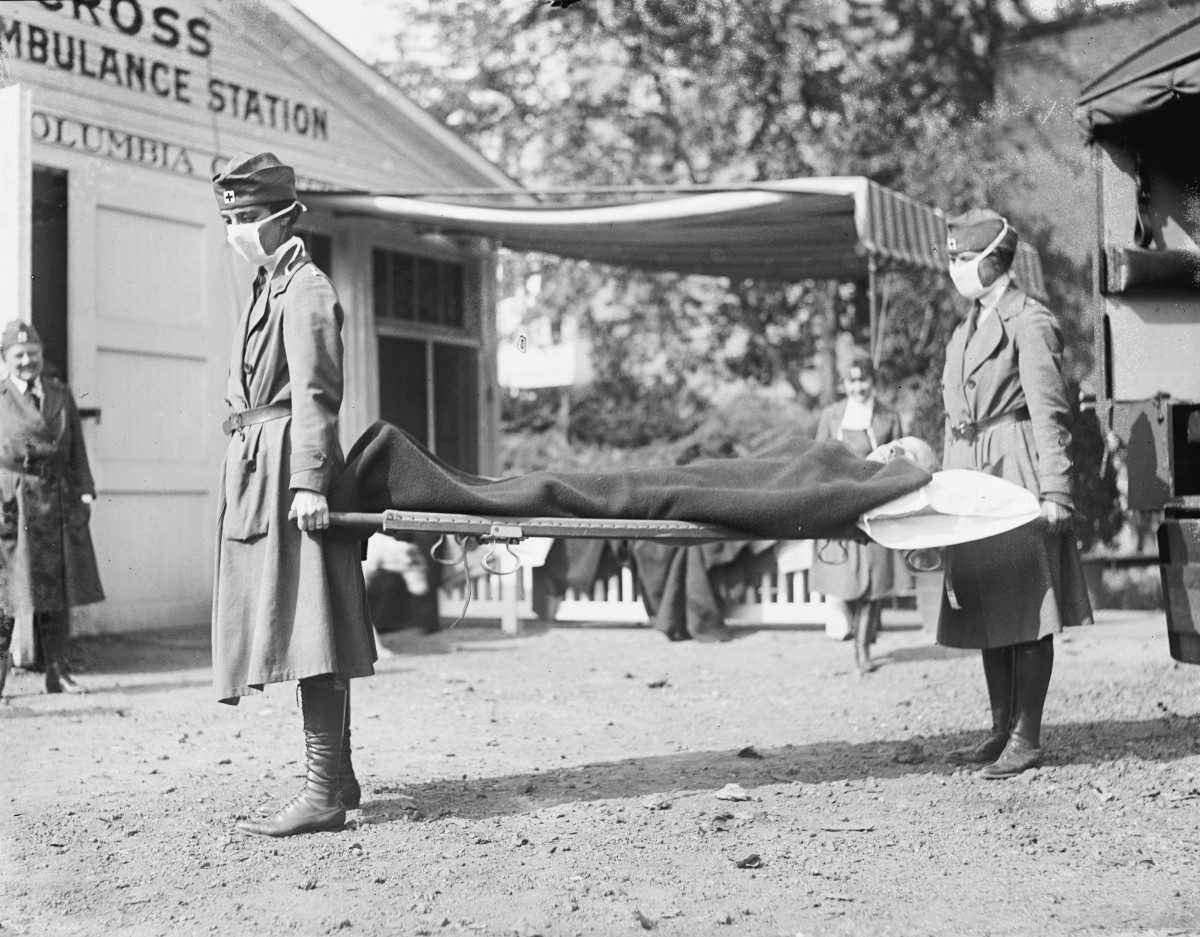
Demonstration at the Red Cross Emergency Ambulance Station in Washington, D.C., during the influenza pandemic of 1918. // Photo: Library of Congress
U of M researchers decode the Spanish Flu, advance in microbial ‘arms race’
The 1918 “Spanish Flu” was one of the most devastating pandemics in human history, killing 50-100 million people. To this day influenza is a major health concern, having killed another 50-100 million people since 1918. The most recent pandemic was the H1N1 flu outbreak in 2009.
“Unlike some other noteworthy diseases, like Smallpox, the flu can infect many different animals including marine life, aquatic birds and other land animals,” said Kevin Coombs, a professor of medical microbiology in the Max Rady College of Medicine at the University of Manitoba. “Therefore, there is little chance of eradicating the virus and the virus’ genetic plasticity allows the flu to rapidly mutate to evade vaccines and anti-virals.”
A new study, published in EBioMedicine by lead authors Coombs and Dr. Darwyn Kobasa from the Public Health Agency of Canada (PHAC), and co-author Dr. Charlene Ranadheera, PHAC, succeeded in measuring thousands of dysregulated cellular proteins. They noticed that many proteins not identified in earlier studies were affected by the Spanish Flu.
Using state-of-the-art mass spectrometry facilities in the Manitoba Centre for Proteomics and Systems Biology at the University of Manitoba to measure upregulation and downregulation of thousands of cellular proteins after virus infection, they compared how the 1918 virus activated specific cellular pathways.
The research team, funded by Canadian Institutes of Health Research and PHAC, found that the Spanish Flu, unlike other flu viruses, induced specific and unique changes early in infection in a master cellular regulator, the Akt/mTOR pathway.
This, Coombs says, suggests that the Spanish Flu had an unusual requirement for this cellular pathway and points the way to more focused studies to prevent future flu pandemics.
“The important differences could explain how it (Spanish Flu) interacts with the cells it infects. This interaction is what made the Spanish Flu so deadly,” Coombs said. “Better understanding host cell responses to the flu may be the only way to stay ahead in the microbial ‘arms race’ in order to prevent re-emergence of this and of other deadly flu viruses.”
Research at the University of Manitoba is partially supported by funding from the Government of Canada Research Support Fund.